The tiger pufferfish, Takifugu
rubripes (fugu) was proposed as a genomic model because of its compact
genome. The genome of fugu was sequenced to the draft level in 2002 as
the second vertebrate genome to be fully sequenced. However, the
assembly of the draft genome sequence is highly fragmented due to the
lack of a chromosomal coordinate. To extend the contiguity of the
assembly to the chromosomal level, we have generated the second
generation genetic map of fugu and anchored the scaffolds of the
assembly to the 22 chromosomes of fugu. The map consists of 1,220
microsatellite markers that provide anchor points to scaffolds covering
86% of the genome assembly.The consolidated genome map of fugu is a
valuable resource for comparative genomics and for elucidating the
genetic basis of the phenotypic diversity of ∼25 species of fugu and
its related species that diverged within the last 5 million years.
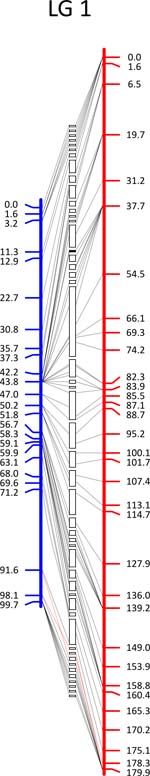
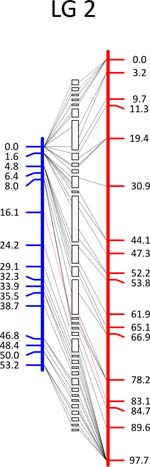
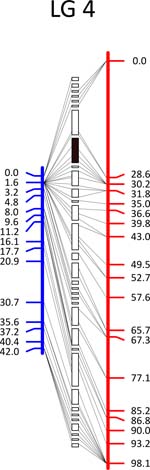
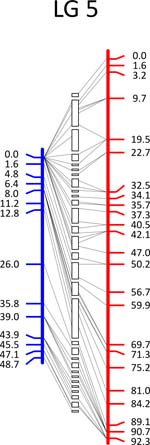
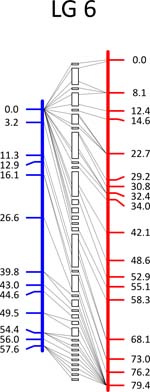
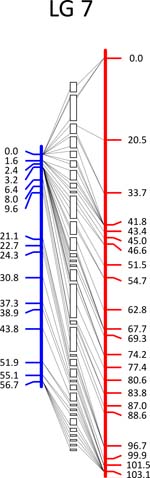
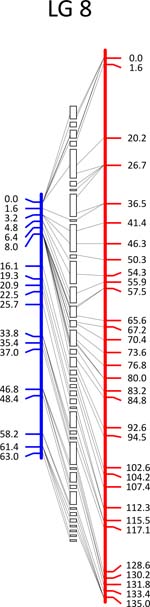
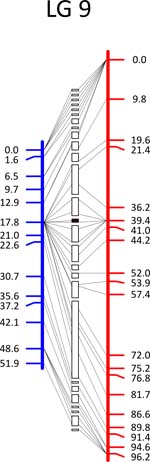
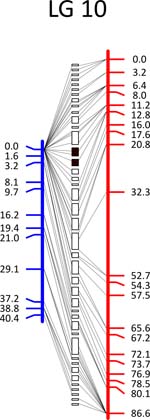
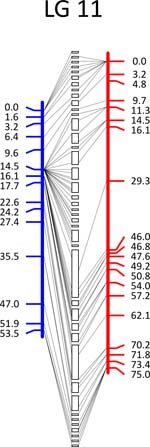
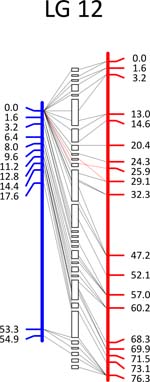
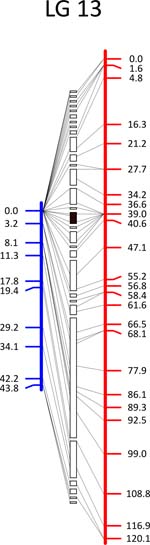
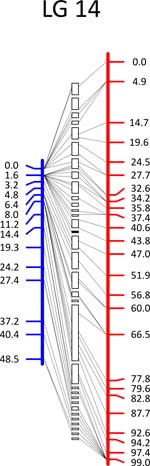
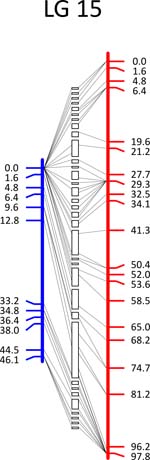
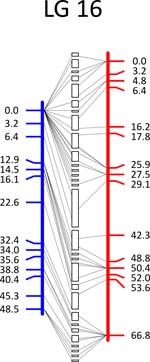
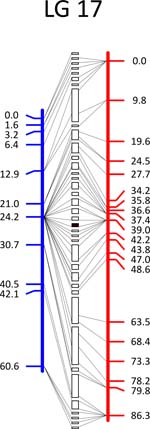
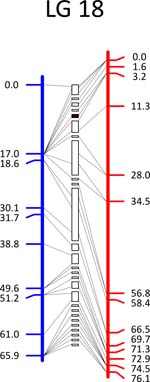
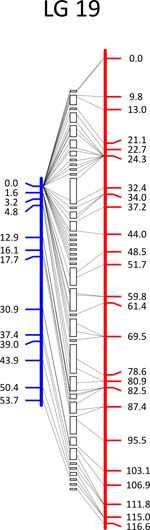
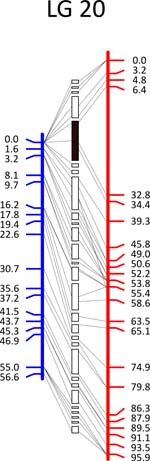
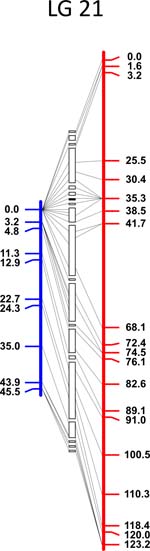
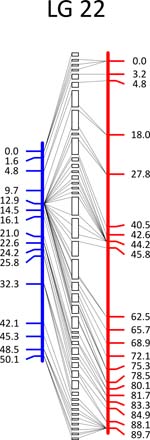
Click each
LG for details.
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |